INL researchers improve detection of pathogenic bacteria with a faster and more specific method
October 2, 2019
Zoonosis or zoonotic diseases happen when bacteria, viruses or parasites allow transmission from animals to humans and are a major concern, both to health authorities and the food industry.
INL Food Quality and Safety Group just released an article showing it is possible to improve the detection of these bacteria responsible for millions of infected people around the globe, with a heavy cost for health systems.
***
Due to the critical impact that foodborne zoonotic diseases have on public health, the detection of pathogenic bacteria in food and water is an important issue for the food industry, for control authorities, and for researchers all over the world. In the European Union (EU), over 320,000 human cases of food-borne zoonotic diseases are reported each year, but the real number is likely to be much higher (WHO 2014; EFSA & ECDC 2018).
To identify any bacterial strains in food, control authorities and the food industry currently use either standard microbiological methods, which take several days or DNA analysis, which provide faster and specific identification, enabling officials to take prompt emergency measures to protect the public when necessary. However, one of the major limitations of the DNA analysis currently used to detect bacteria is that these do not discriminate between live and dead bacteria present. Thus, if using, for instance, a Polymerase Chain Reaction, known as PCR, to detect bacteria, there may be many false-positive results due to the detection of dead bacteria which do not represent any hazard for the consumers. This is a major issue for the producers, as these type of false-positive results will cause a delay in the release of the products and additional costs associated with longer storage periods, additional analysis, etc.
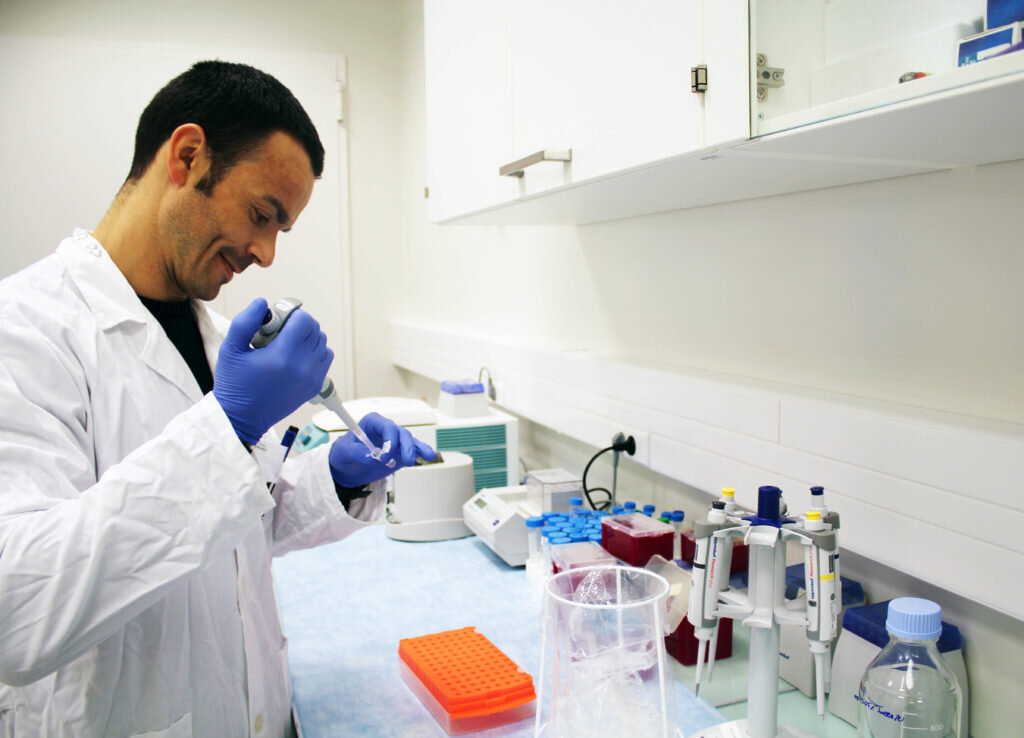
At INL, the Food Quality and Safety (FQS) Research Group has developed and evaluated a methodology for more rapid and specific detection of bacteria, especially for one which is responsible for roughly 50% of the cases of salmonellosis in Europe, the viable Salmonella Enteritidis. Their work was published this year in Food Control, https://www.sciencedirect.com/science/article/pii/S0956713518306388*.
The novelty of their methodology lies in the combination of a phage amplification assay with real-time PCR. Bacteriophages are viruses that infect bacteria. They are highly host-specific, meaning that all phages do not infect all bacterial species. This is an added value as can be used as an indirect way to assess the presence of certain bacteria species.
Their method was tested in chicken samples (one of the major sources of salmonellosis), and proved to provide reliable results in just 10 hours! This has a considerable advantage when compared to either conventional methods which can take up to 1 week, or current DNA-based methods which although take approx. 24-48 hours, still pose the risk of false positives associated with the presence of DNA from dead microorganisms.
By using bacteriophages, their method detects only live bacteria, since these viruses do not replicate in dead microorganisms, thus assuring that a positive result is directly linked to the presence of live bacteria.
“As these viruses actually kill the bacteria that they infect, we first let the Salmonella to recover and start multiplying, then we add a known concentration of bacteriophages, and finally by using real-time PCR we can track the changes in viral DNA”, explains Alejandro Garrido-Maestu, a staff researcher in the FQS group at INL and co-author of the study. In this sense, he states that “the fact that the viruses multiply inside the bacteria, allows to improve the sensitivity of the methodology and achieve a significant time reduction in the assay. An increase in viral DNA will indicate that the viruses are multiplying, and so that the pathogen is present and alive; on the other hand, if there is no change in the DNA concentration, then we can assure that the bacteria are either absent or if present, they are dead and do not represent any health risk for the consumer.”